Calculate pi amino acid in a polypeptide1/28/2024 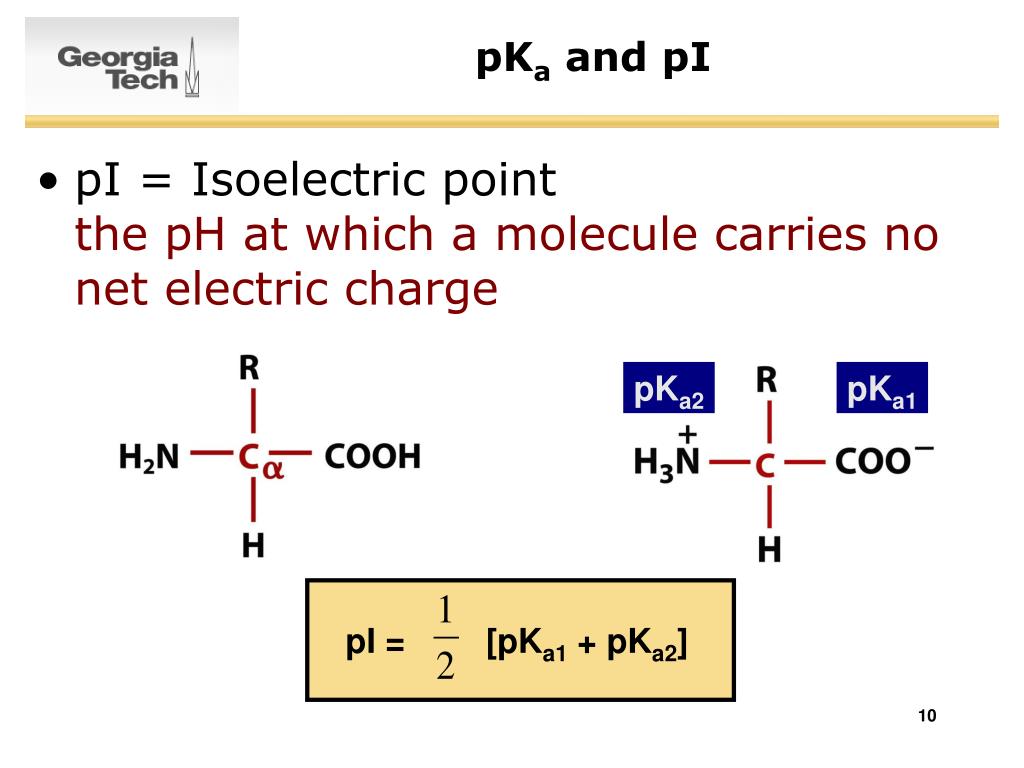  
Sequences and have yielded information important for comparative genomics, as well as experimental applications. Predictions of the complete assemblage of proteins an organism is capable of expressing have been made from annotated genome Our analysesĪlso suggest that the proportions of proteomes consisting of membrane-associated proteins may be currently underestimated. Of proteomes and are associated with the need for different pI values depending on subcellular localization. Of whole proteome pI values in bacteria and archaea and the trimodal character in eukaryotes are likely to be general properties On the basis of this association between pI and subcellular localization, we conclude that the bimodal character Furthermore, nuclear proteins have a broader distribution that may account for the third mode observed To cytosolic and integral membrane proteins have pI distributions that appear to correspond with the two observed modes ofīacteria and archaea. Generated from the full database by selecting on subcellular localization shows that sequences annotated as corresponding A similar analysis on subsets of a nonredundant protein sequence database 1999) and reveal a trimodality in eukaryotic genomes. Of pI values confirm the bimodality that has been observed previously for bacterial and archaeal genomes ( Van Bogelen et al. This article analyzesĭistributions of pI values estimated computationally for all predicted ORFs in a selection of fully sequenced genomes. Isoelectric point (pI) values have long been a standard measure for distinguishing between proteins.
0 Comments
Leave a Reply.AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
